SBEToolbox 1.3.3
:: DESCRIPTION
SBEToolbox (Systems Biology and Evolution Toolbox) is being developed in MATLAB as a menu-driven GUI software to determine various statistics of the biological network. Some of its features include (but not limited to) algorithms to create random networks (small-world, ring lattice etc..), deduce clusters in the network (MCL, mCode, clusterOne), compute various network topology measures etc…
::DEVELOPER
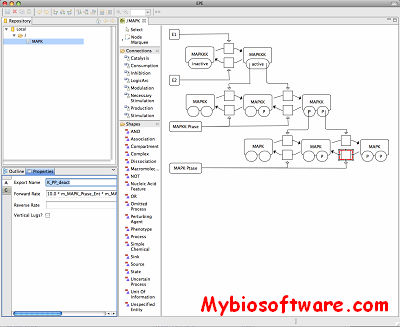
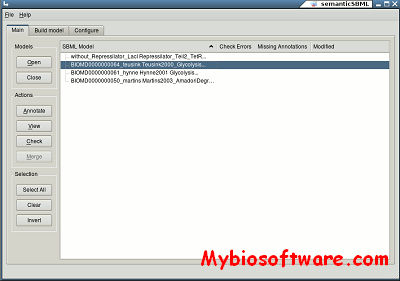
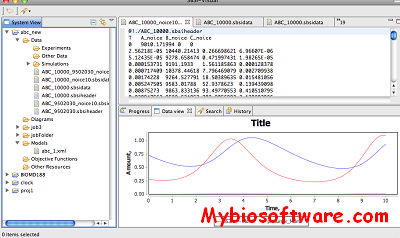
:: SCREENSHOTS
N/A
:: REQUIREMENTS
- Linux / Windows / MacOsX
- MATLAB
:: DOWNLOAD
:: MORE INFORMATION
Citation
Evol Bioinform Online. 2013 Sep 1;9:355-62. doi: 10.4137/EBO.S12012. eCollection 2013.
SBEToolbox: A Matlab Toolbox for Biological Network Analysis.
Konganti K1, Wang G, Yang E, Cai JJ.