SemanticSBML 2.0
:: DESCRIPTION
SemanticSBML create, check, annotate, merge SBML (System Biology Markup Lanugage) models. The program includes a graphical user interface as well as a console interface that can process batch jobs.
Semantic annotations can be used to define – among other things – the biochemical meaning of model elements. The tool semanticSBML is made to handle such annotations. It helps you to edit them and to check and merge your models.
::DEVELOPER
semanticSBML team at Humboldt University Berlin
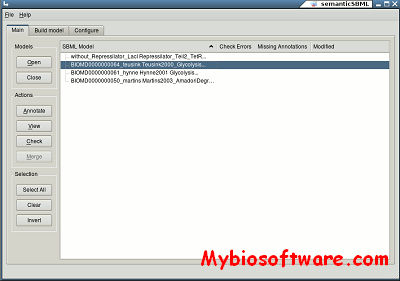
:: SCREENSHOTS
:: REQUIREMENTS
:: DOWNLOAD
:: MORE INFORMATION
Citation
For citing semanticSBML, please refer to: Krause F, Uhlendorf J., Lubitz T., Schulz M., Klipp E., Liebermeister W. (2010),
Annotation and merging of SBML models with semanticSBML,
Bioinformatics 26 (3), 421-422, doi:10.1093/bioinformatics/btp642.