SeaView 5.0.5
:: DESCRIPTION
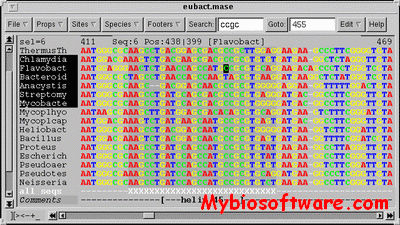
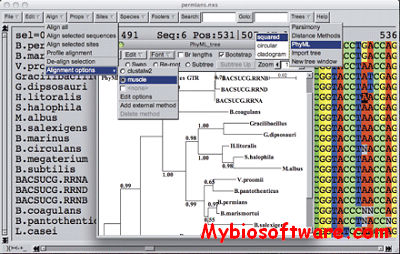
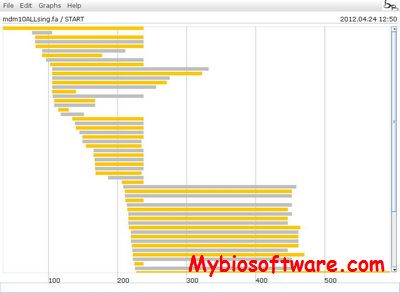
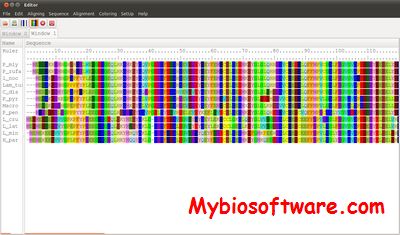
SeaView is a multiplatform, graphical user interface for multiple sequence alignment and molecular phylogeny.
::DEVELOPER
:: SCREENSHOTS
:: REQUIREMENTS
- Windows / Mac OsX / Linux
:: DOWNLOAD
:: MORE INFORMATION
Citation
Gouy M., Guindon S. & Gascuel O. (2010)
SeaView version 4 : a multiplatform graphical user interface for sequence alignment and phylogenetic tree building.
Molecular Biology and Evolution 27(2):221-224.