MPAlign-editor 3.0.0
:: DESCRIPTION
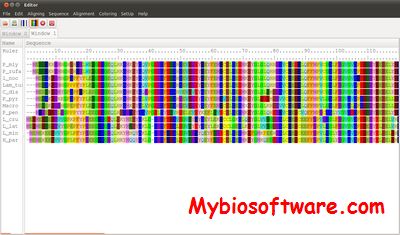
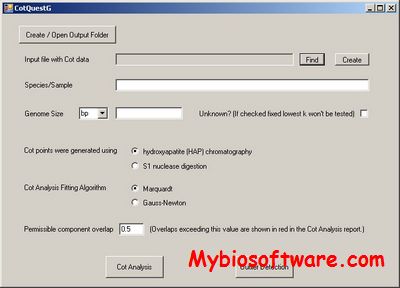
MPAlign-editor is a graphical tool for DNA and Protein sequences and alignments.
::DEVELOPER
Frederico Arnoldi (fgcarnoldi@gmail.com)
:: SCREENSHOTS
:: REQUIREMENTS
- Linux / Windows / Mac OsX
- Python
:: DOWNLOAD
:: MORE INFORMATION
Citation
MPAlign editor: uma ferramenta gráfica e intuitiva para alinhamentos moleculares.
Frederico GC Arnoldi 2005
Proceedings of II Workshop TIDIA—FAPESP