ArgusLab 4.0.1
:: DESCRIPTION
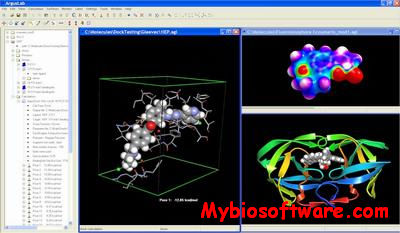
ArgusLab is a molecular modeling, graphics, and drug design program.The ArgusDock docking engine, implemented in ArgusLab4.0, approximates an exhaustive search method, with similarities to DOCK and Glide. Flexible ligand docking is possible with ArgusLab, where the ligand is described as a torsion tree and grids are constructed that overlay the binding site. Ligand’s root node (group of bonded atoms that do not have rotatable bonds) is placed on a search point in the binding site and a set of diverse and energetically favorable rotations is created. For each rotation, torsions in breadth-first order are constructed and those poses that survive the torsion search are scored. The N-lowest energy poses are retained and the final set of poses undergoes coarse minimization, re-clustering and ranking.
::DEVELOPER
Mark A. Thompson , Planaria Software LLC, Seattle, WA
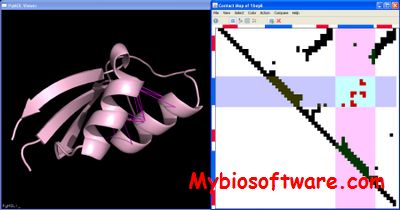
:: SCREENSHOTS
:: REQUIREMENTS
- Windows
:: DOWNLOAD
:: MORE INFORMATION
Citation
Thompson, M. A. (2004).
Molecular docking using ArgusLab, an efficient shape-based search algorithm and the AScore scoring function.
ACS meeting, Philadelphia, 172, CINF 42, PA.