xGDBvm
:: DESCRIPTION
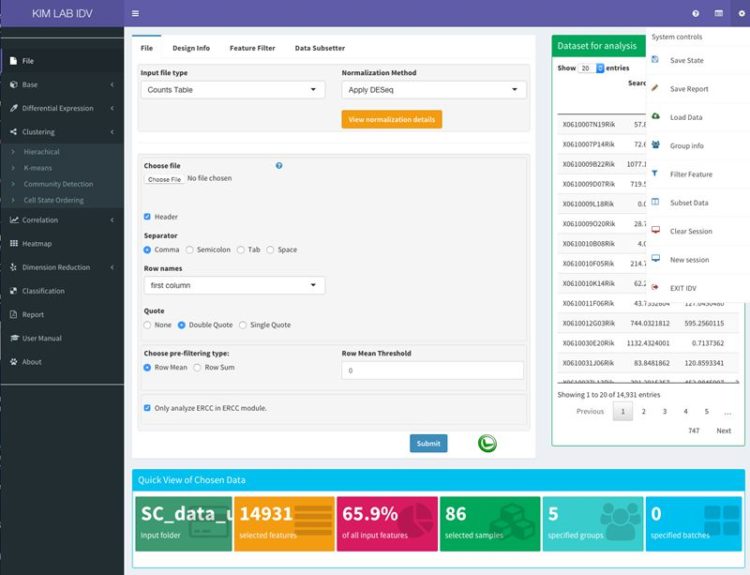
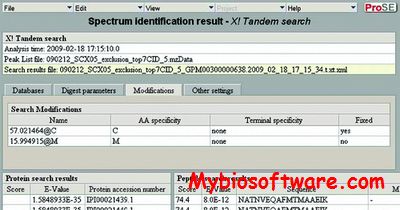
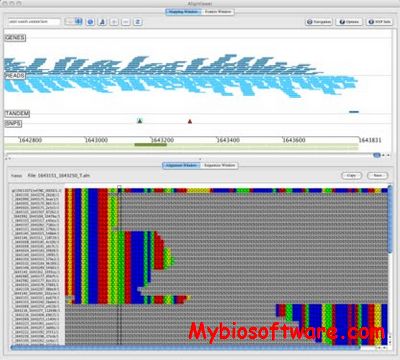
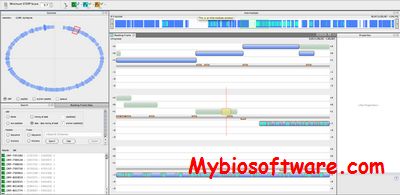
xGDBvm is a web-based platform for annotating and visualizing eukaryotic genomes.
::DEVELOPER
The Brendel Group @ Indiana University
:: REQUIREMENTS
- Linux
- PHP
:: DOWNLOAD
:: MORE INFORMATION
Citation
Duvick J, Standage DS, Merchant N, Brendel VP.
xGDBvm: A Web GUI-Driven Workflow for Annotating Eukaryotic Genomes in the Cloud.
Plant Cell. 2016 Apr;28(4):840-54. doi: 10.1105/tpc.15.00933. Epub 2016 Mar 28. PMID: 27020957; PMCID: PMC4863385.