BiQ Analyzer 3.0 / BiQAnalyzer HT / BiQ Analyzer HiMod
:: DESCRIPTION
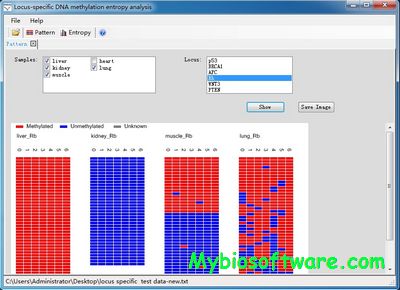
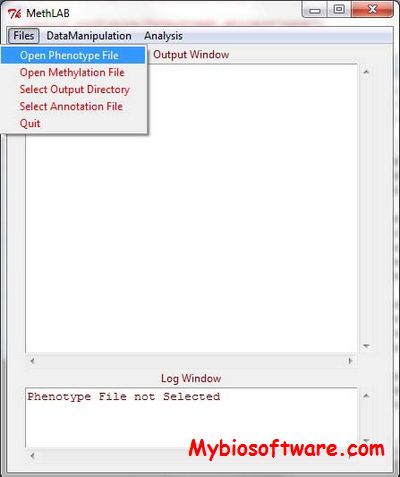
BiQ Analyzer is a software tool for easy visualization and quality control of DNA methylation data from bisulfite sequencing.
BiQAnalyzer HT is the major upgrade of the popular BiQAnalyzer tool specifically designed to aid the analysis of high-throughput bisulfite amplicon sequencing data.
BiQ Analyzer HiMod is the major upgrade of the popular BiQAnalyzer HT tool specifically designed to aid the analysis of high-throughput bisulfite amplicon sequencing data.
::DEVELOPER
Max-Planck-Institut Informatik
:: SCREENSHOTS
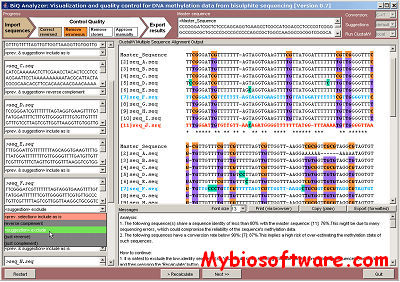
:: REQUIREMENTS
- Windows / MacOsX / Linux
- Java
:: DOWNLOAD
 BiQ Analyzer ,BiQ Analyzer HT ,
BiQ Analyzer ,BiQ Analyzer HT ,
:: MORE INFORMATION
Citation
Bock, C., S. Reither, T. Mikeska, M. Paulsen, J. Walter and T. Lengauer (2005).
“BiQ Analyzer: visualization and quality control for DNA methylation data from bisulfite sequencing.”
Bioinformatics 21(21): 4067-8.
Pavlo Lutsik, Lars Feuerbach, Julia Arand, Thomas Lengauer, Jörn Walter and Christoph Bock
BiQ Analyzer HT: locus-specific analysis of DNA methylation by high-throughput bisulfite sequencing.
Nucl. Acids Res. (2011) 39 (suppl 2): W551-W556.
BiQ Analyzer HiMod: an interactive software tool for high-throughput locus-specific analysis of 5-methylcytosine and its oxidized derivatives.
Becker D, Lutsik P, Ebert P, Bock C, Lengauer T, Walter J.
Nucleic Acids Res. 2014 Jul;42(Web Server issue):W501-7. doi: 10.1093/nar/gku457.