SCLpredT
:: DESCRIPTION
SCLpredT is an enhanced version of SCLpred (Subcellular Localisation), in that: it incorporates homology information to proteins of known localization ; it is trained on a larger dataset; in has one more output class (“organelle”).
::DEVELOPER
:: SCREENSHOTS
N/A
:: REQUIREMENTS
- Web Browser
:: DOWNLOAD
 NO
NO
:: MORE INFORMATION
Citation
Springerplus. 2013 Oct 3;2:502. doi: 10.1186/2193-1801-2-502. eCollection 2013.
SCLpredT: Ab initio and homology-based prediction of subcellular localization by N-to-1 neural networks.
Adelfio A1, Volpato V, Pollastri G.
TAGOPSIN 1.3 – Collating Taxa-specific Gene and Protein Functional and Structural Information
TAGOPSIN 1.3
:: DESCRIPTION
TAGOPSIN (TAxonomy, Gene, Ontology, Protein, Structure INtegrated) retrieves select data from NCBI Taxonomy, NCBI Nucleotide, UniProtKB, Gene Ontology, Pfam, EBI SIFTS and RCSB PDB, and assembles them in the database management system PostgreSQL. TAGOPSIN is an organism-centred data warehousing tool that works with prokaryotic and eukaryotic organisms as well as viruses.
::DEVELOPER
:: SCREENSHOTS
N/A
:: REQUIREMENTS
- Linux / MacOsX
- Java
- PostgreSQL
:: DOWNLOAD
:: MORE INFORMATION
Citation
Bundhoo E, Ghoorah AW, Jaufeerally-Fakim Y.
TAGOPSIN: collating taxa-specific gene and protein functional and structural information.
BMC Bioinformatics. 2021 Oct 23;22(1):517. doi: 10.1186/s12859-021-04429-5. PMID: 34688246; PMCID: PMC8541804.
ANAT 3.0 – Inference and Analysis of Functional Networks of Proteins
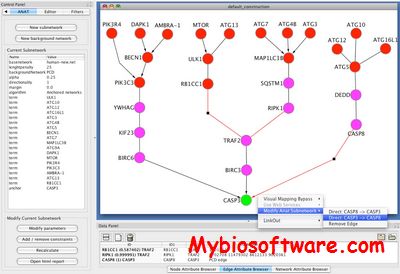
ANAT 3.0
:: DESCRIPTION
ANAT (Advanced Network Analysis Tool) , is an all-in-one resource that provides access to up-to-date large-scale physical association data in several organisms, advanced algorithms for network reconstruction, and a number of tools for exploring and evaluating the obtained network models
::DEVELOPER
:: SCREENSHOTS
:: REQUIREMENTS
:: DOWNLOAD
:: MORE INFORMATION
Citation
Signorini LF, Almozlino T, Sharan R.
ANAT 3.0: a framework for elucidating functional protein subnetworks using graph-theoretic and machine learning approaches.
BMC Bioinformatics. 2021 Oct 27;22(1):526. doi: 10.1186/s12859-021-04449-1. PMID: 34706638.
ANAT: A Tool for Constructing and Analyzing Functional Protein Networks.
N. Yosef, E. Zalckvar, A. D. Rubinstein, M. Homilius, N. Atias, L. Vardi, I. Berman, H. Zur, A. Kimchi, E. Ruppin and R. Sharan
Sci. Signal. 4, pl1 (2011).
Erebus – Protein Substructure Search Server
Erebus
:: DESCRIPTION
Erebus is a web server that searches the entire Protein Data Bank for a given substructure defined by a set of atoms of interest, such as the binding scaffolds for small molecules.
::DEVELOPER
:: SCREENSHOTS
N/A
:: REQUIREMENTS
- Web Browser
:: DOWNLOAD
 NO
NO
:: MORE INFORMATION
Citation
Bioinformatics. 2011 May 1;27(9):1327-9. doi: 10.1093/bioinformatics/btr129. Epub 2011 Apr 1.
Rigid substructure search.
Shirvanyants D1, Alexandrova AN, Dokholyan NV.
INTERSPIA – INTER-Species Protein Interaction Analysis
INTERSPIA
:: DESCRIPTION
INTERSPIA is a web-based application for the analysis and visualization of the dynamics of protein-protein interactions among multiple species.
::DEVELOPER
Bioinformatics Laboratory, Konkuk University
:: SCREENSHOTS
N/A
:: REQUIREMENTS
- Web browser
:: DOWNLOAD
 NO
NO
:: MORE INFORMATION
Citation
INTERSPIA: a web application for exploring the dynamics of protein-protein interactions among multiple species.
Kwon D, Lee D, Kim J, Lee J, Sim M, Kim J.
Nucleic Acids Res. 2018 Jul 2;46(W1):W89-W94. doi: 10.1093/nar/gky378.
PSTP-finder 0.4.3.1 – Protein Surface Transient Pocket Finder
PSTP-finder 0.4.3.1
:: DESCRIPTION
PSTP-finder is a software developed in order to analyse a molecular dynamic of a protein structure and highlight time ranges in which a transient pocket could be found around certain aminoacid residues on the protein surface.
::DEVELOPER
SBL (Stuctural Biology Laboratory), University of Siena
:: SCREENSHOTS
N/A
:: REQUIREMENTS
- Linux
- C Compiler
:: DOWNLOAD
:: MORE INFORMATION
Sadic / ProCoCoA 1.1b10 – Simple Atom Depth Index Calculator / Protein Core Composition Analyzer
Sadic / ProCoCoA 1.1b10
:: DESCRIPTION
Sadic/ProCoCoA is able to calculate the atom depth value for every atom of a protein structure, giving a lot of intrinsic information of 3D composition of the macromolecule and its core.
::DEVELOPER
SBL (Stuctural Biology Laboratory), University of Siena
:: SCREENSHOTS
N/A
:: REQUIREMENTS
- Web browser
:: DOWNLOAD
 NO
NO
:: MORE INFORMATION
Citation:
Comput Biol Chem. 2013 Apr;43:29-34. doi: 10.1016/j.compbiolchem.2012.12.007. Epub 2012 Dec 31.
ProCoCoA: A quantitative approach for analyzing protein core composition.
Bottini S1, Bernini A, De Chiara M, Garlaschelli D, Spiga O, Dioguardi M, Vannuccini E, Tramontano A, Niccolai N.
DISCO 1.0 – Structure Determination of Protein Homo-oligomers
DISCO 1.0
:: DESCRIPTION
DISCO is software to perform structure determination of protein homo-oligomers with cyclic symmetry.DISCO computes oligomeric protein structures using geometric constraints derived from RDCs and intermolecular distance restraints such as NOEs or disulfide bonds. When a reliable subunit structure can be calculated from intramolecular restraints, DISCO guarantees that all satisfying oligomer structures will be discovered, yet can run in minutes to hours on only a single desktop-class computer.
::DEVELOPER
Donald Lab at Duke University
:: SCREENSHOTS
N/A
:: REQUIREMENTS
- Windows / Linux / MacOSX
- Java
:: DOWNLOAD
:: MORE INFORMATION
Citation
Jeffrey W. Martin, Anthony K. Yan, Chris Bailey-Kellogg, Pei Zhou, and Bruce R. Donald.
A graphical method for analyzing distance restraints using residual dipolar couplings for structure determination of symmetric protein homo-oligomers.
Protein Science, 20(6):970-985, 2011.
pydca 1.0 – Direct Coupling analysis software for Protein and RNA sequences
pydca 1.0
:: DESCRIPTION
pydca is Python implementation of direct coupling analysis (DCA) of residue coevolution for protein and RNA sequence families using the mean-field and pseudolikelihood maximization algorithms.
::DEVELOPER
Multiscale Biomolecular Simulation
:: SCREENSHOTS
n/a
:: REQUIREMENTS
- Linux
- Python
- C++
:: DOWNLOAD
:: MORE INFORMATION
Citation
Zerihun MB, Pucci F, Peter EK, Schug A.
pydca v1.0: a comprehensive software for direct coupling analysis of RNA and protein sequences.
Bioinformatics. 2020 Apr 1;36(7):2264-2265. doi: 10.1093/bioinformatics/btz892. PMID: 31778142.
Customprodbj v1.2.0 – Customized Protein Database Construction
Customprodbj v1.2.0
:: DESCRIPTION
Customprodbj is a Java-based tool for customized protein database construction. It can be used to (1) build a customized database based on single VCF file, (2) build a customized database based on multiple VCF files from a sample and (3) build a customized database based on multiple VCF files from multiple samples.
::DEVELOPER
:: SCREENSHOTS
n/a
:: REQUIREMENTS
- Linux / Windows
- Java
:: DOWNLOAD
:: MORE INFORMATION