MUMmer 4.0.0Beta2/ pymummer 0.11.0
:: DESCRIPTION
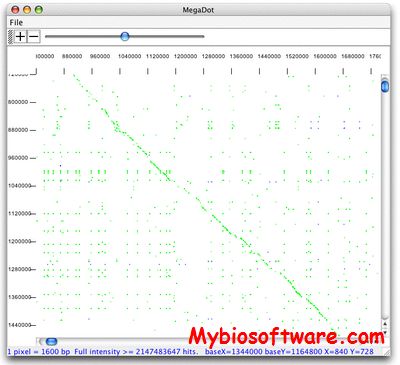
MUMmer is a system for rapidly aligning entire genomes, whether in complete or draft form.MUMmer can also align incomplete genomes; it can easily handle the 100s or 1000s of contigs from a shotgun sequencing project, and will align them to another set of contigs or a genome using the NUCmer program included with the system. If the species are too divergent for a DNA sequence alignment to detect similarity, then the PROmer program can generate alignments based upon the six-frame translations of both input sequences.
pymummer is Python3 module for running MUMmer and reading the output
::DEVELOPER
MUMmer team
:: SCREENSHOTS
N/A
:: REQUIREMENTS
:: DOWNLOAD
 MUMmer, pymummer
MUMmer, pymummer
:: MORE INFORMATION
MUMmer is now an open source project.
Citation:
MUMmer4: A fast and versatile genome alignment system
G. Marçais , A.L. Delcher, A.M. Phillippy, R. Coston, S.L. Salzberg, A. Zimin,
PLoS computational biology (2018), 14(1): e1005944.