GOfact 20051017
:: DESCRIPTION
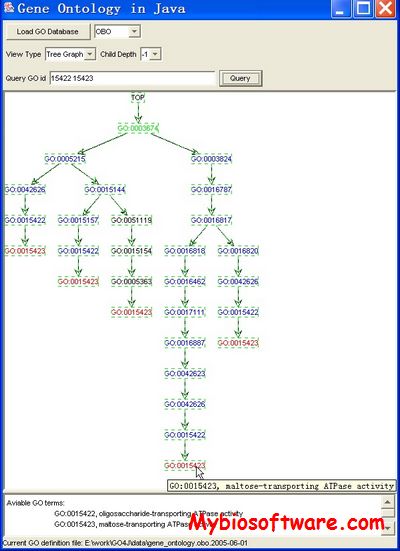
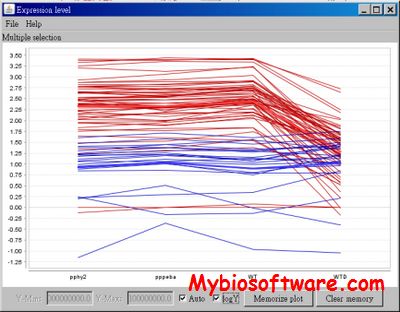
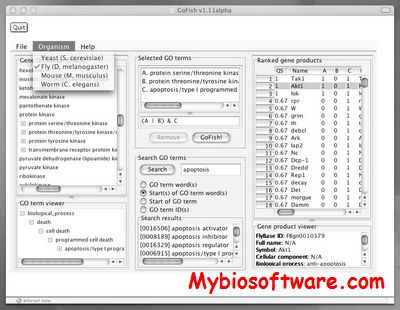
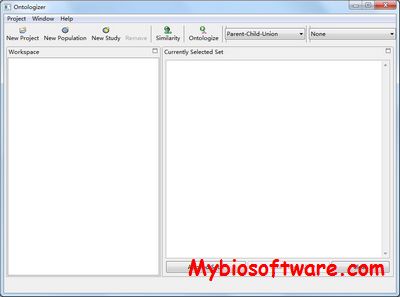
GOfact is developed to identify the functional distribution and the significantly enriched functional categories of the proteomic expression profile. It would be helpful for understanding the overall functions of these identified proteins and supply the fundamental information for further bioinformatics exploration.
::DEVELOPER
:: SCREENSHOTS
N/A
:: REQUIREMENTS
- Linux/Windows
- Perl
:: DOWNLOAD
:: MORE INFORMATION
Citation
Li Dong, Li Jian-Qi, Ouyang Shu-Guang, Wang Jian, Xu Xiaojie, Zhu Yun-Ping, He Fu-Chu.
An Integrated Strategy for Functional Analysis in Large-scale Proteomic Research by Gene Ontology.
Biochemistry and Biophysics 2005,32(11):1026-1029