Cluster 3.0 20190830
:: DESCRIPTION
Cluster 3.0 is an enhanced version of Cluster, which was originally developed by Michael Eisen while at Stanford University.
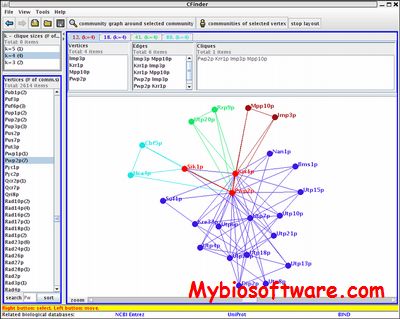
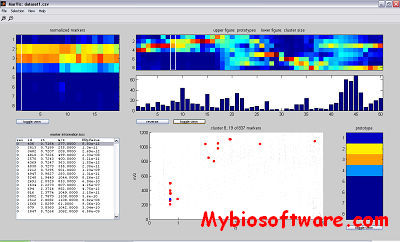
Cluster is program that provide a computational and graphical environment for analyzing data from DNA microarray experiments, or other genomic datasets. The program Cluster can organize and analyze the data in a number of different ways.
The main improvement consists of the k-means algorithm, which now includes multiple trials to find the best clustering solution. This is crucial for the k-means algorithm to be reliable.The routine for self-organizing maps was extended to include 2D rectangular geometries. The Euclidean distance and the city-block distance were added to the available measures of similarity.
::DEVELOPER
Michiel de Hoon of the University of Tokyo
:: SCREENSHOTS
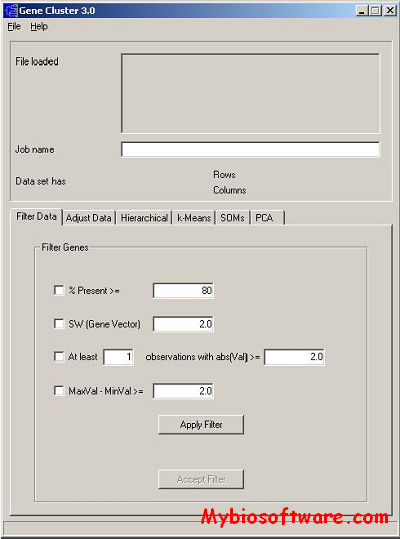

:: REQUIREMENTS
- Windows
- Mac OS X
- Linux/Unix with Motif
:: DOWNLOAD
 Cluster
Cluster
:: MORE INFORMATION
Reference: M. J. L. de Hoon, S. Imoto, J. Nolan, and S. Miyano:
Open Source Clustering Software.
Bioinformatics, 20 (9): 1453–1454 (2004).