PIVOT 2.0
:: DESCRIPTION
PIVOT (Protein Interactions VisualizatiOn Tool) is a Java-based tool, for visualizing protein-protein interactions. It is rich in features that help the users navigate and interpret the interactions map, as well as graph-theory algorithms for easily connecting remote proteins to the displayed map.PIVOT is not limited to the data present in any database, but instead allows the users to create their own dataset, by importing any available lists of interactions and combining them together. PIVOT is currently configured to work with proteins from four different species (human, yeast, drosophila and mouse), and present functional annotations, designations of homologs from the four species, and links to information pages. The protein data is stored in an MS-Access file, easily modifiable by the users to enter their own data.
::DEVELOPER
Ron Shamir’s lab
:: SCREENSHOTS
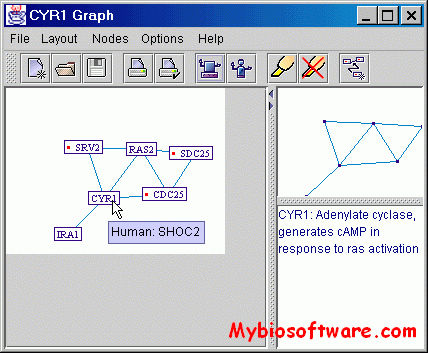
:: REQUIREMENTS
- Windows / Linux / MacOSX
- Java
:: DOWNLOAD
 PIVOT
PIVOT
:: MORE INFORMATION
Citation
Bioinformatics. 2004 Feb 12;20(3):424-5.
PIVOT: protein interacions visualizatiOn tool.
Orlev N1, Shamir R, Shiloh Y.


 NO
NO