SBW 2.12.2
:: DESCRIPTION
SBW ( Systems Biology Workbench), is an open source framework connecting heterogeneous software applications.
Researchers in quantitative systems biology make use of a large number of different software packages for modeling, analysis, visualization, and general data manipulation. The Systems Biology Workbench (SBW), is a software framework that allows heterogeneous application components-written in diverse programming languages and running on different platforms-to communicate and use each others’ capabilities via a fast binary encoded-message system.
SBW goal was to create a simple, high performance, open-source software infrastructure which is easy to implement and understand. SBW enables applications (potentially running on separate, distributed computers) to communicate via a simple network protocol. The interfaces to the system are encapsulated in client-side libraries that we provide for different programming languages.
::DEVELOPER
Sauro Lab at University of Washington
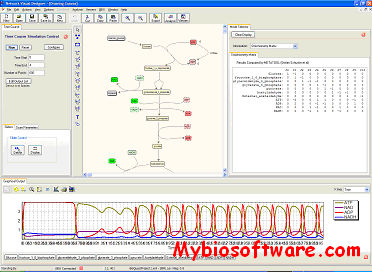
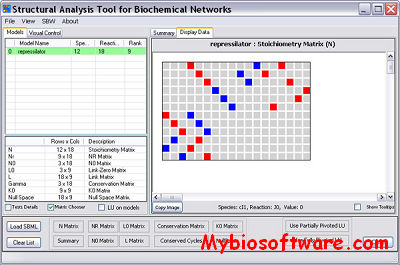
:: SCREENSHOTS
:: REQUIREMENTS
- Windows/MacOsX/Linux
:: DOWNLOAD
:: MORE INFORMATION
Citation
Sauro, Hucka, Finney, Wellock, Bolouri, Doyle, Kitano.
“Next generation simulation tools: the Systems Biology Workbench and BioSPICE integration.”
OMICS. 2003 Winter;7(4):355-72.