DREM 2.0.4/ mirDREM
:: DESCRIPTION
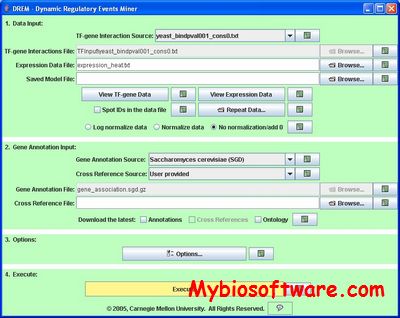
The DREM (Dynamic Regulatory Events Miner) allows one to model, analyze, and visualize transcriptional gene regulation dynamics. The method of DREM takes as input time series gene expression data and static or dynamic transcription factor-gene interaction data (e.g. ChIP-chip data), and produces as output a dynamic regulatory map. The dynamic regulatory map highlights major bifurcation events in the time series expression data and transcription factors potentially responsible for them.
mirDREM supports the use of microRNAs in DREM
::DEVELOPER
Systems Biology Group – Carnegie Mellon University
:: SCREENSHOTS
:: REQUIREMENTS
- Linux / MacOsX / Windows
- Java
:: DOWNLOAD
:: MORE INFORMATION
Citation
DREM 2.0: Improved reconstruction of dynamic regulatory networks from time-series expression data.
Schulz MH, Devanny WE, Gitter A, Zhong S, Ernst J, Bar-Joseph Z.
BMC Syst Biol. 2012 Aug 16;6(1):104.
MH Schulz, KV Pandit, CLL Cardenas, A Namasivayam, N Kaminsky and Z. Bar-Joseph.
Reconstructing dynamic miRNA regulated interaction networks
PNAS, August 28, 2013, doi: 10.1073/pnas.1303236110