EDISA 1.0
:: DESCRIPTION
EDISA (Extended Dimension Iterative Signature Algorithm) is a novel probabilistic clustering approach for 3D gene-condition-time datasets. Based on mathematical definitions of gene expression modules, the EDISA samples initial modules from the dataset which are then refined by removing genes and conditions until they comply with the module definition. A subsequent extension step ensures gene and condition maximality. We applied the algorithm to a synthetic dataset and were able to successfully recover the implanted modules over a range of background noise intensities.
::DEVELOPER
the Center for Bioinformatics Tübingen (Zentrum für Bioinformatik Tübingen, ZBIT).
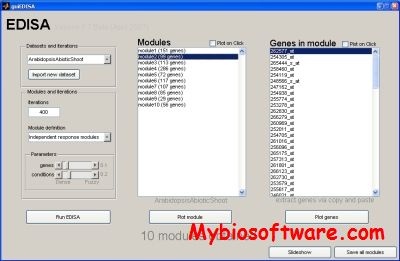
:: SCREENSHOTS
:: REQUIREMENTS
- Linux/ WIndows
- Matlab
:: DOWNLOAD
:: MORE INFORMATION
Citation
Jochen Supper, Martin Strauch, Dierk Wanke, Klaus Harter, Andreas Zell:
EDISA: extracting biclusters from multiple time-series of gene expression profiles
BMC Bioinformatics 2007, 8:334