PhosphoLogo
:: DESCRIPTION
PhosphoLogo is a tool for generating information-based sequence logos from a list of equal-length peptide sequences centered on the phosphorylated residues. It has been designed to meet the needs of analyzing protein kinase phospho-motifs, but can be used for other modifications or unmodified peptides.
::DEVELOPER
Epithelial Systems Biology Laboratory
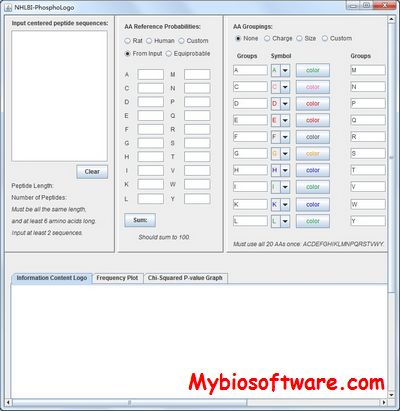
:: SCREENSHOTS
:: REQUIREMENTS
- Windows/ Linux/ MacOsX
- JRE
:: DOWNLOAD
:: MORE INFORMATION
Citation
Am J Physiol Cell Physiol. 2012 Oct 1;303(7):C715-27. doi: 10.1152/ajpcell.00166.2012. Epub 2012 Jun 20.
Identifying protein kinase target preferences using mass spectrometry.
Douglass J1, Gunaratne R, Bradford D, Saeed F, Hoffert JD, Steinbach PJ, Knepper MA, Pisitkun T.