PathwayExplorer 3.0.2
:: DESCRIPTION
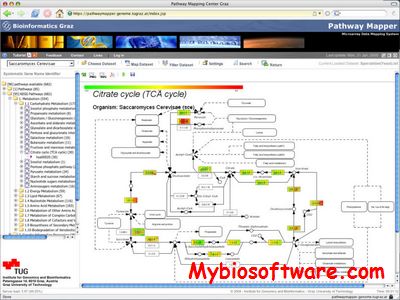
The PathwayExplorer is an easy to use and platform independent interactive Java drawing tool for the construction of biological pathway diagrams in a visual way and the annotation of the components and interactions between them. This novel drawing tool integrates the possibilities of charting elements with different attributes (size, color, labels), drawing connections between elements in distinct characteristics (color, structure, width, arrows), as well as adding links to molecular biology databases, promoter sequences, information on the function of the genes or gene products, and references. The result of the editing process is a PNG (portable network graphics) file for the images and XML (extended markup language) file for the appropriate links. Further it is possible to map expression datasets to the pathways by using gene RefSeq ids. It is also possible to use a SOAP interface to access the pathways online.
::DEVELOPER
Genomics & Bioinformatics Graz, Graz University of Technology
:: SCREENSHOTS
:: REQUIREMENTS
- Linux / Windows / MacOsX
- Java
- Oracle / PostgreSQL / MySQL
:: DOWNLOAD
:: MORE INFORMATION
Citation
Mlecnik B, Scheideler M, Hackl H, Hartler J, Sanchez-Cabo F, Trajanoski Z.
PathwayExplorer: web service for visualizing high-throughput expression data on biological pathways.
Nucleic Acids Res. 33: W633-W637 (2005)