Gcorr
:: DESCRIPTION
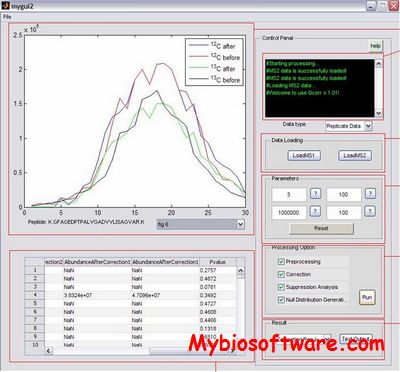
Gcorr corrects peptide profiles that satisfy correction conditions, and it can predict fold change null distributions at different intensity levels. Subsequently, the significance P-values of measured fold changes can be estimated based on the predicted null distributions. We have used an 1:1 LC-FTMS label-free dataset pair collected based on the same sample to verify that our predicted null distributions conforms to that of the observed null distribution.
::DEVELOPER
:: SCREENSHOTS
::REQUIREMENTS
- Linux/ Windows/MacOsX
- MatLab
:: DOWNLOAD
:: MORE INFORMATION
Citation
Proteome Sci. 2011 Oct 14;9 Suppl 1:S2. doi: 10.1186/1477-5956-9-S1-S2.
Processing methods for signal suppression of FTMS data.
Ma X1, Cui J, Zhang J.