UniMoG
:: DESCRIPTION
UniMoG (former DCJ) is a software tool unifying five genome rearrangement distance models: double cut and join (DCJ), restricted DCJ, Hannenhalli and Pevzner (HP), inversion only and translocation only. It allows computing all of these five distances between pairs of genomes represented as sequences of oriented common blocks.
DCJ (double-cut-and-join) computes the double-cut-and-join distance between two genomes and an optimal sorting scenario that transforms one genome into the other. It operates on the most general model of genomes with a mixed collection of linear and circular chromosomes. The sorting process includes all classical rearrangement operations such as inversions, translocations, transpositions, block interchanges, fusions and fissions.
::DEVELOPER
Rafael Friesen, Julia Mixtacki, Jens Stoye
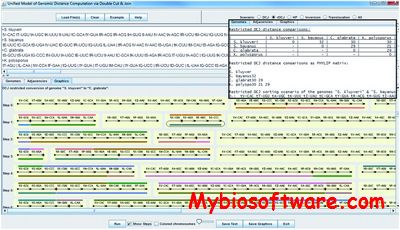
:: SCREENSHOTS
:: REQUIREMENTS
- Linux / Mac / Windows
- Java
:: DOWNLOAD
:: MORE INFORMATION
Citation
Bioinformatics. 2012 Oct 1;28(19):2509-11. Epub 2012 Jul 18.
UniMoG–a unifying framework for genomic distance calculation and sorting based on DCJ.
Hilker R1, Sickinger C, Pedersen CN, Stoye J.
A. Bergeron, J. Mixtacki, J. Stoye.
A unifying view of genome rearrangements
Proceedings of WABI 2006, LNBI 4175, 163-173, 2006.