JAMIE 0.91
:: DESCRIPTION
JAMIE (Joint Analysis of Multiple IP Experiments) is a R package to perform the joint analysis. The genome is assumed to consist of background and potential binding regions (PBRs). PBRs have context-dependent probabilities to become bona fide binding sites in individual datasets. This model captures the correlation among datasets, which provides basis for sharing information across experiments. Real data tests illustrate the advantage of JAMIE over a strategy that analyzes individual datasets separately.
::DEVELOPER
Ji Lab
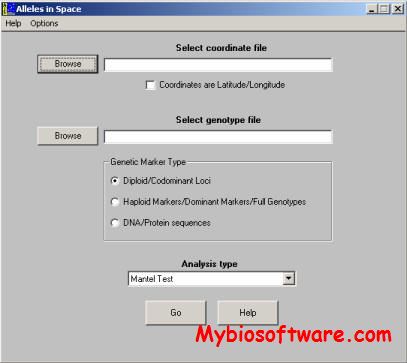
:: SCREENSHOTS
N/A
:: REQUIREMENTS
- Windows / MacOsX / Linux
- R Package
- affy and affxparser
:: DOWNLOAD
:: MORE INFORMATION
Citation
Wu H. and Ji H. (2010)
JAMIE: Joint analysis of multiple ChIP-chip experiments.
Bioinformatics (2010) 26 (15): 1864-1870.