CSS-Palm 4.0
:: DESCRIPTION
CSS-Palm is a computer program for palmitoylation site prediction, Clustering and Scoring Strategy for Palmitoylation Sites Prediction.The program’s prediction performance is encouraging with highly positive Jack-Knife validation results (sensitivity 82.16% and specificity 83.17% for cut-off score 2.6).
::DEVELOPER
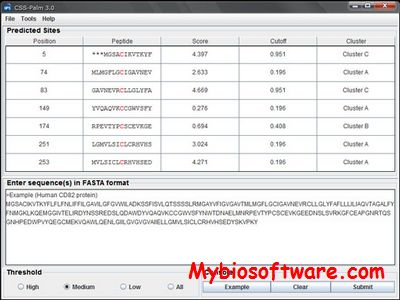
:: SCREENSHOTS
:: REQUIREMENTS
- WIndows / Linux / MacOsX
- Java
:: DOWNLOAD
:: MORE INFORMATION
Citation
CSS-Palm 2.0: an updated software for palmitoylation sites prediction
Jian Ren, Longping Wen, Xinjiao Gao, Changjiang Jin, Yu Xue and Xuebiao Yao.
Protein Engineering, Design and Selection.2008 21(11):639-644