UNAFold 4.0 / mfold 3.6 / MFold Interface 1.18
:: DESCRIPTION
UNAFold (Unified Nucleic Acid Folding & Hybridization Package) is an integrated collection of programs that simulate folding, hybridization, and melting pathways for one or two single-stranded nucleic acid sequences. Folding (secondary structure) prediction for single-stranded RNA or DNA combines free energy minimization, partition function calculations and stochastic sampling. For melting simulations, the package computes entire melting profiles, not just melting temperatures. UV absorbance at 260 nm, heat capacity change (C(p)), and mole fractions of different molecular species are computed as a function of temperature.
mfold has been replaced by UNAFold.Although UNAFold will install without mfold_util, the sir_graph and boxplot_ng programs from the mfold_util package are required in order to obtain structure plots and dot plots from UNAFold. Install mfold_util before installing UNAFold. Versions 3.4 and higher of mfold contains all of the non-interactive programs in mfold_util, so a separate download is not required.
MFold Interface is Perl/Tk GUI Interface for Dr. Michael Zuker’s MFold.
::DEVELOPER
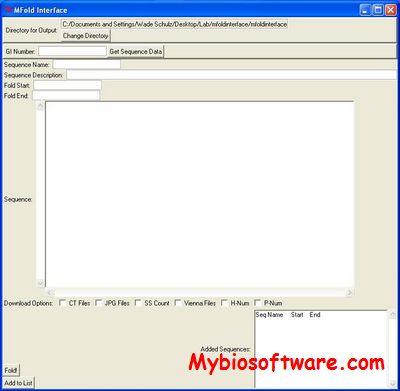
:: SCREENSHOTS
:: REQUIREMENTS
- Linux / Windows / Mac OsX
- glut/OpenGL
- GD Library
- gnuplot
- perl
:: DOWNLOAD
:: MORE INFORMATION
Citation
Methods Mol Biol. 2008;453:3-31.
UNAFold: software for nucleic acid folding and hybridization.
Markham NR, Zuker M.