Sybil 1.5.1
:: DESCRIPTION
Sybil is a web-based software package for comparative genomics, whose primary goal is to facilitate the analysis and visualization of comparative genome data, with a particular emphasis on protein and gene cluster data. Herein, a two-phase protein clustering algorithm, used to generate protein clusters suitable for analysis through Sybil and a method for creating graphical displays of protein or gene clusters that span multiple genomes are described. When combined, these two relatively simple techniques provide the user of the Sybil software (The Institute for Genomic Research [TIGR] Bioinformatics Department) with a browsable graphical display of his or her “input” genomes, showing which genes are conserved based on the parameters supplied to the protein clustering algorithm. For any given protein cluster the graphical display consists of a local alignment of the genomes in which the clustered genes are located. The genomes are arranged in a vertical stack, as in a multiple alignment, and shaded areas are used to connect genes in the same cluster, thus displaying conservation at the protein level in the context of the underlying genomic sequences. The authors have found this display-and slight variants thereof-useful for a variety of annotation and comparison tasks, ranging from identifying “missed” gene models or single-exon discrepancies between orthologous genes, to finding large or small regions of conserved gene synteny, and investigating the properties of the breakpoints between such regions.
::DEVELOPER
J. Craig Venter Institute
:: SCREENSHOTS
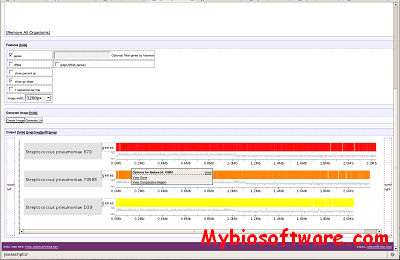

:: REQUIREMENTS
:: DOWNLOAD
 Sybil
Sybil
:: MORE INFORMATION
Citation:
Crabtree J, Angiuoli SV, Wortman JR, White OR.
Sybil: methods and software for multiple genome comparison and visualization.
Methods Mol Biol. 2007;408:93-108.


