RNASeqBrowser V3.1
:: DESCRIPTION
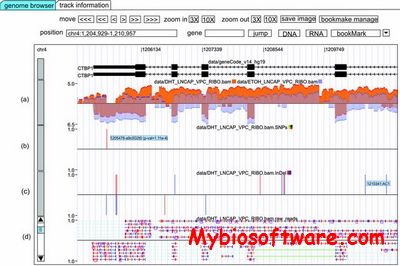
RNASeqBrowser is a visualization tool that incorporates and extends the functionality of the UCSC genome browser. For example, RNASeqBrowser simultaneously displays read coverage, SNPs, InDels and raw read tracks with other BED and wiggle tracks — all being dynamically built from the BAM file.
::DEVELOPER
:: SCREENSHOTS
:: REQUIREMENTS
- Windows / Linux / MacOsX
- java
:: DOWNLOAD
:: MORE INFORMATION