DomainGraph 3.01
:: DESCRIPTION
DomainGraph is the successor tool of DomainNetworkBuilder and works as Java plugin for Cytoscape, a free open-source software platform for visualization and analysis of biomolecular networks. This plugin decomposes protein networks into domain-domain interactions and generates a new network of interacting domains. It also allows the integration of exon expression data measured using Affymetrix GeneChip microarrays, which supports the analysis of alternative splicing events and the characterization of their effects on protein and domain interaction networks.
::DEVELOPER
Max-Planck-Institut Informatik
:: SCREENSHOTS
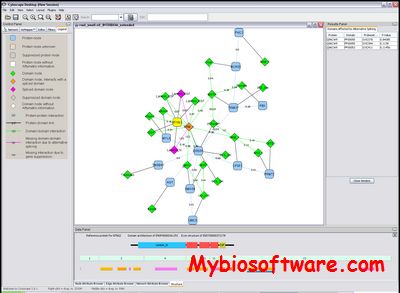
:: REQUIREMENTS
:: DOWNLOAD
 DomainGraph
DomainGraph
:: MORE INFORMATION
Citation:
Emig, D., Salomonis, N., Baumbach, J., Lengauer, T., Conklin, BR., Albrecht, M. (2010)
AltAnalyze and DomainGraph: analyzing and visualizing exon expression data.
Nucleic Acids Res. 2010 July 1; 38