NNN 1.01
:: DESCRIPTION
NNN (Nearest Neighbor Networks) is a graph-based algorithm to generate clusters of genes with similar expression profiles. This method produces clusters based on overlapping cliques within an interaction network generated from mutual nearest neighborhoods. This focus on nearest neighbors rather than on absolute distance measures allows us to capture clusters with high connectivity even when they are spatially separated, and requiring mutual nearest neighbors allows genes with no sufficiently similar partners to remain unclustered. We compared the clusters generated by NNN with those generated by eight other clustering methods. NNN was particularly successful at generating functionally coherent clusters with high precision, and these clusters generally represented a much broader selection of biological processes than those recovered by other methods.
::DEVELOPER
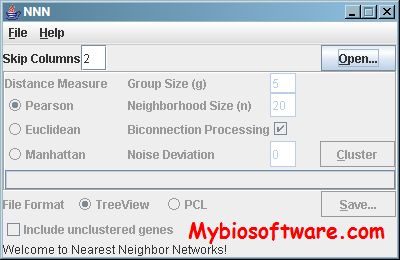
:: SCREENSHOTS
:: REQUIREMENTS
- Windows / Linux / Mac OsX
- java
:: DOWNLOAD
:: MORE INFORMATION
Citation
Huttenhower, C., Flamholz, A., Landis, J., Sahi, S., Myers, C., Olszewski, K., Hibbs, M., Siemers, N., Troyanskaya, O., Coller, H.,
“Nearest Neighbor Networks: Clustering Expression Data Based on Gene Neighborhoods“,
BMC Bioinformatics 8:250, 2007