Topali 2.5 r3
:: DESCRIPTION
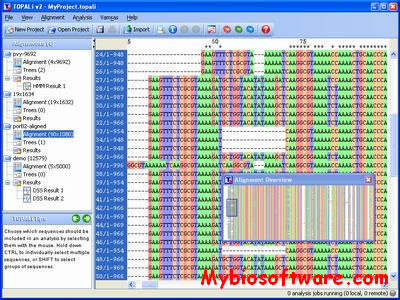
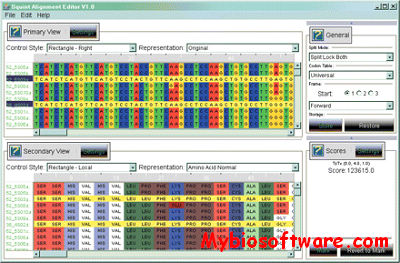
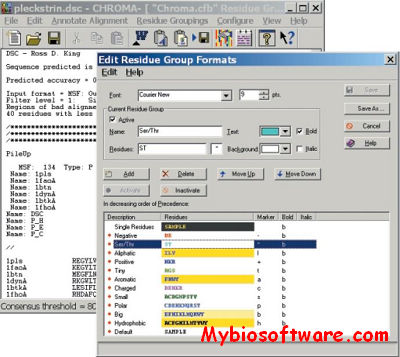
Topali (tree TOPology-related analysis of ALignments Interface) is a software for statistical and evolutionary analysis of multiple sequence alignments.The extended TOPALi v2 provides phylogenetic model selection, Bayesian analysis (BA) and Maximum Likelihood (ML) phylogenetic tree estimation, detection of sites under positive selection, and recombination breakpoint location analysis.
::DEVELOPER
Information & Computational Sciences, The James Hutton Institute
:: SCREENSHOTS
:: REQUIREMENTS
- Windows/ Linux / Mac OsX
:: DOWNLOAD
:: MORE INFORMATION
Citation
Milne I, Lindner D, Bayer M, Husmeier D, McGuire G, Marshall DF and Wright F (2008),
TOPALi v2: a rich graphical interface for evolutionary analyses of multiple alignments on HPC clusters and multi-core desktops,
Bioinformatics 25 (1), 126-127.