VISCOE 1.0
:: DESCRIPTION
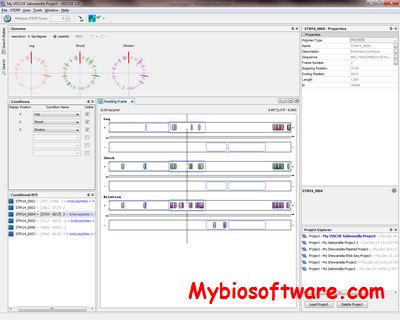
VISCOE is a genomic integration tool to simultaneously visualize proteomics experiments across multiple conditions. VISCOE maps peptides onto proteins on the appropriate DNA strand and offers peptide- and protein-centric searches to find regions of qualitative differences (up to six conditions). VISCOE is also capable of integration of RNA-seq transcriptomic data for simultaneous visualization. RNA-seq data can be represented in SAM or WIG format.
::DEVELOPER
Computational Biology & Bioinformatics ,Pacific Northwest National Laboratory
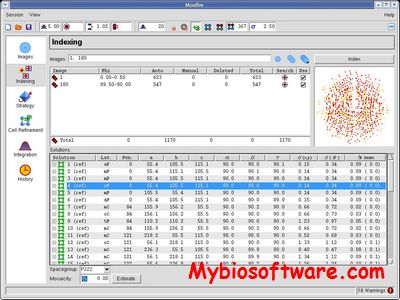
:: SCREENSHOTS
:: REQUIREMENTS
- Windows / Linux / MacOsX
- Java
:: DOWNLOAD
:: MORE INFORMATION