Paxtools 5.1.0 / PaxtoolsR 1.26.0
:: DESCRIPTION
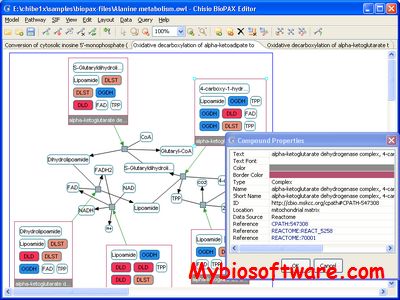
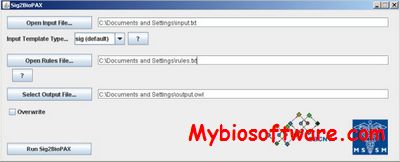
Paxtools is a Java library specially designed for accessing and manipulating data in BioPAX format. The Paxtools Java programming library for BioPAX has been developed to help software developers readily support the import, export and validation of BioPAX-formatted data for various uses in their software . Using Paxtools, a range of BioPAX-compatible software has been developed, including browsers, visualizers, querying engines, editors and converters
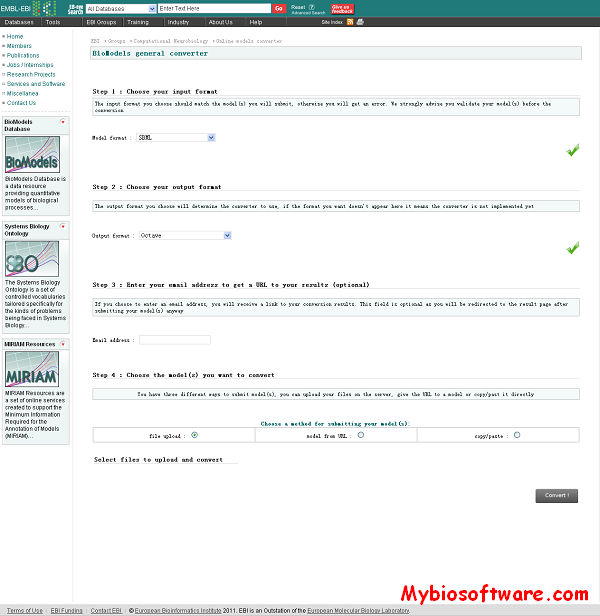
PaxtoolsR provides a set of R functions for interacting with BioPAX OWL files using Paxtools and the querying Pathway Commons (PC) molecular interaction database that are hosted by the Computational Biology Center at Memorial Sloan-Kettering Cancer Center (MSKCC).
::DEVELOPER
Computational Biology Center at MSKCC
:: SCREENSHOTS
N/A
:: REQUIREMENTS
- Windows / Linux / MacOSX
- Java / R/ BioConductor
:: DOWNLOAD
 Paxtools , PaxtoolsR
Paxtools , PaxtoolsR
:: MORE INFORMATION
Citation:
PaxtoolsR: Pathway Analysis in R Using Pathway Commons.
Luna A, Babur Ö, Aksoy BA, Demir E, Sander C.
Bioinformatics. 2015 Dec 18. pii: btv733.
Demir et al.
The BioPAX community standard for pathway data sharing
Nature Biotechnology 28 , 935–942 (2010) doi:10.1038/nbt.1666