SpliceSeq 2.1
:: DESCRIPTION
SpliceSeq provides a quick, easy method of investigating alternative mRNA splicing in next generation mRNA sequence data. The tool may be used on a single mRNA-Seq sample to identify genes with multiple spliceforms or on a pair of samples to identify differential splicing between the samples. Sequence reads are mapped to splice graphs that unambiguously quantify the inclusion level of each exon and splice junction. The graphs are then traversed to predict the protein isoforms that are likely to result from the observed exon and splice junction reads. UniProt annotations are mapped to each protein isoform to identify potential functional impacts of alternative splicing.
::DEVELOPER
Department of Bioinformatics and Computational Biology, The University of Texas MD Anderson Cancer Center
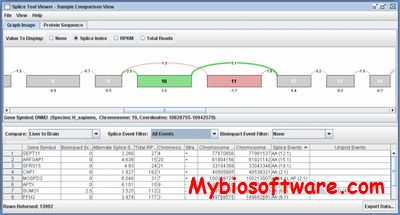
:: SCREENSHOTS
:: REQUIREMENTS
- Linux/Windows/MacOsX
- Java
:: DOWNLOAD
:: MORE INFORMATION
Citation
Bioinformatics. 2012 Sep 15;28(18):2385-7. Epub 2012 Jul 20.
SpliceSeq: a resource for analysis and visualization of RNA-Seq data on alternative splicing and its functional impacts.
Ryan MC, Cleland J, Kim R, Wong WC, Weinstein JN.