SBMLsqueezer 2.1
:: DESCRIPTION
SBMLsqueezer generates kinetic equations for biochemical networks according to context of each reaction. When used as a plugin for CellDesigner it uses the information from the SBGN representation of all network components. In the stand-alone mode, SBMLsqueezer evaluates the Systems Biology Ontology (SBO) annotations to extract this information.
The rate laws that can be produced by SBMLsqueezer include several types of generalized mass action; detailed and generalized enzyme kinetics, various types of Hill equations, S- and H-systems, and additive models for gene regulation. User defined settings specify which equation to apply for any type of reaction and how to ensure unit consistency of the model. Equations can be created using contextual menus. All newly created parameters are equipped with the derived unit and annotated with SBO terms if available and meaningful textual names. MathML is inserted directly into the SBML file. LaTeX or text export of ordinary differential equations is provided.
::DEVELOPER
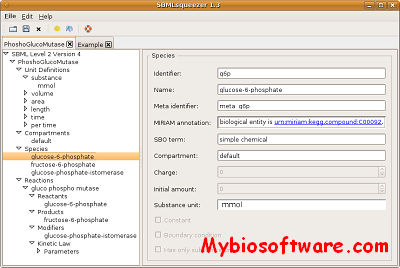
:: SCREENSHOTS
:: REQUIREMENTS
- Windows / Linux / Mac OsX
- Java Runtime Environment
- CellDesigner (Plugin mode)
:: DOWNLOAD
:: MORE INFORMATION
Citation
SBMLsqueezer: a CellDesigner plug-in to generate kinetic rate equations for biochemical networks.
Dräger A, Hassis N, Supper J, Schröder A, Zell A.
BMC Syst Biol. 2008 Apr 30;2:39. doi: 10.1186/1752-0509-2-39.