LACK 4.3
:: DESCRIPTION
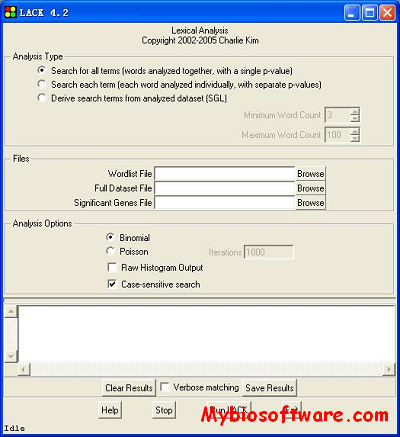
LACK addresses whether or not a theme is actually overrepresented in your significant genes list. The program takes a list of significant genes and a list of user-specified search terms, and counts the number of genes which contain one of the search terms. Then, the program takes a random set of genes of the same size as the significant set from a genome annotation file and counts the hits. This process is repeated a user-specified number of times so that statistics regarding the randomness of the frequencies can be calculated. Statistics and histogram data is output to a text file.
::DEVELOPER
:: SCREENSHOTS
:: REQUIREMENTS
:: DOWNLOAD
 LACK for Win ; Source Code ; Manual ; Sample files (zipped)
LACK for Win ; Source Code ; Manual ; Sample files (zipped)
:: MORE INFORMATION
The software is copyrighted under the terms of the GNU General Public License. You can view this license at http://www.gnu.org/licenses/gpl.txt.
If you use LACK, please cite:
Kim CC, Falkow S.
Significance analysis of lexical bias in microarray data.
BMC Bioinformatics. 2003 Apr 3;4(1):12.