iGEMDOCK 2.1
:: DESCRIPTION
iGEMDOCK is a Graphical Environment for Recognizing Pharmacological Interactions and Virtual Screening.Pharmacological interactions are useful for identifying lead compounds and understanding ligand binding mechanisms for a therapeutic target. Currently, these interactions are often inferred from a set of active compounds that were acquired experimentally. Moreover, most docking programs loosely coupled the stages of structure-based virtual screening (VS) from preparations through to post-screening analysis. An integrated VS environment, which provides the friendly interface to seamlessly combine different-stage programs for VS and identifying the pharmacological interactions from screening compounds, is valuable for drug discovery. Here, we developed an easy-to-use graphic environment, iGEMDOCK, for the docking, virtual screening, and post-screening analysis. For post-screening analysis, iGEMDOCK can enrich the hit rate and provide biological insights by deriving the pharmacological interactions from screening compounds. The pharmacological interactions represent conserved interacting residues that often form binding pockets with specific physico-chemical properties to play the essential functions of the target protein. Experiment results show that the success rate of iGEMDOCK is 78% (root-mean-square derivations below 2.0 angstrom) on 305 protein-compound complexes. For virtual screening, pharmacological interactions derived by iGEMDOCK often involve the biological functions and enrich the hit rates on three public sets (i.e., estrogen receptor α for antagonists (ER) and agonists (ERA) and thymidine kinase (TK)). We believe that iGEMDOCK is useful for understanding the ligand binding mechanisms and discovering lead compounds.
::DEVELOPER
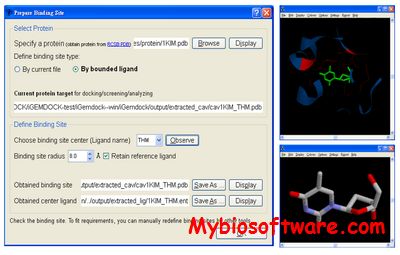
:: SCREENSHOTS
:: REQUIREMENTS
- Linux/Windows
:: DOWNLOAD
:: MORE INFORMATION
Citation
K.-C. Hsu, Y.-F. Chen, S.-R. Lin and J.-M. Yang*,
“iGEMDOCK: A Graphical Environment of enhancing GEMDOCK using pharmacological interactions and postscreening analysis,”
BMC Bioinformatics, 12(Suppl 1):S33, 2011