GSAA 2.0 / GSAA-SNP / GSAA-Seq
:: DESCRIPTION
GSAA (Gene Set Association Analysis) is a computational method that integrates gene expression analysis with genome wide association studies to determine whether an a priori defined sets of genes shows statistically significant, concordant differences with respect to gene expression profiles and genotypes between two biological states. Gene sets are generally a group of genes that are putatively functionally related, co-regulated, or tightly linked on the same chromosome.
GSAA-SNP (Gene Set Association Analysis-SNP) is a computational method that determines whether an a priori defined sets of genes shows statistically significant, concordant differences with respect to genotypes between two biological states. Gene sets are generally a group of genes that are putatively functionally related, co-regulated, or tightly linked on the same chromosome.
GSAA-Seq (Gene Set Association Analysis for RNA-Seq) is a computational method that evaluates whether an a priori defined sets of genes shows statistically significant, concordant differences with respect to RNA-Seq gene expression profiles between two biological states. Gene sets are generally a group of genes that are putatively functionally related, co-regulated, or tightly linked on the same chromosome.
::DEVELOPER
Furey Lab at The University of North Carolina at Chapel Hill and Mukherjee Lab at Duke University.
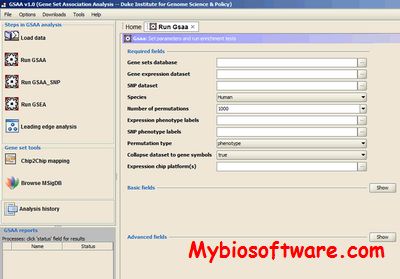
:: SCREENSHOTS
:: REQUIREMENTS
- Windows/Linux/MacOsX
- Java
:: DOWNLOAD
:: MORE INFORMATION
Citation
GSAASeqSP: a toolset for gene set association analysis of RNA-Seq data.
Xiong Q, Mukherjee S, Furey TS.
Sci Rep. 2014 Sep 12;4:6347. doi: 10.1038/srep06347.
Genome Res. 2012 Feb;22(2):386-97. doi: 10.1101/gr.124370.111. Epub 2011 Sep 22.
Integrating genetic and gene expression evidence into genome-wide association analysis of gene sets.
Xiong Q, Ancona N, Hauser ER, Mukherjee S, Furey TS.