FootPrinter 2.1
:: DESCRIPTION
FootPrinter is a program for phylogenetic footprinting. Phylogenetic footprinting is a method that identifies putative regulatory elements in DNA sequences. It identifies regions of DNA that are unusually well conserved across a set of orthologous sequences.
MicroFootPrinter is a front end to the FootPrinter phylogenetic footprinting program, but with specific focus on prokaryotic genomes. You supply a prokaryotic species and gene of interest, and you set a few parameters (or leave them at their default values). MicroFootPrinter will then find related prokaryotes containing a homologous gene, and run FootPrinter to identify motifs that are well conserved across the cis-regulatory regions of these homologous genes.
::DEVELOPER
The Computational & Synthetic Biology group – COMPUTER SCIENCE & ENGINEERING at UNIVERSITY OF WASHINGTON
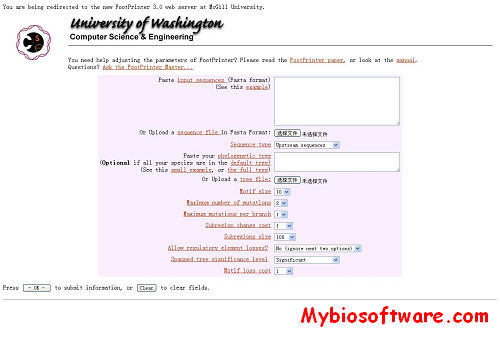
:: SCREENSHOTS
FootPrinter
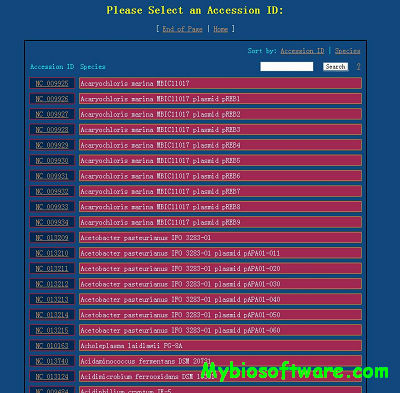
MicroFootPrinter
:: REQUIREMENTS
- Linux
- GCC
:: DOWNLOAD
 FootPrinter Source Code ; Manual
FootPrinter Source Code ; Manual
:: MORE INFORMATION
Citation for FootPrinter:
- M. Blanchette, B. Schwikowski, M. Tompa, “Algorithms for phylogenetic footprinting”, J. Comput. Biol., vol. 9 (2002) 211-23. Pubmed 12015878.
- M. Blanchette, M. Tompa, “Discovery of regulatory elements by a computational method for phylogenetic footprinting”, Genome Res., vol. 12 (2002) 739-48.Pubmed 11997340. Supplement.
- M. Blanchette, M. Tompa, “FootPrinter: A program designed for phylogenetic footprinting”, Nucleic Acids Res., vol. 31 (2003) 3840-2. Pubmed 12824433.
Citation for MicroFootPrinter:
- Neph, S. and Tompa, M., MicroFootPrinter: a Tool for Phylogenetic Footprinting in Prokaryotic Genomes. Nucleic Acids Research, vol. 34, July 2006, W366-W368.