Cytoscape 3.8.2
:: DESCRIPTION
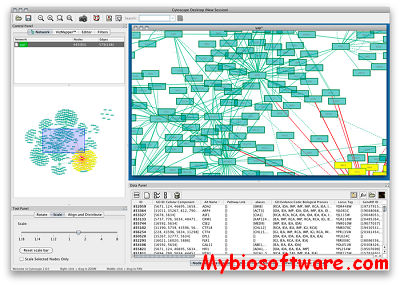
Cytoscape is an open source bioinformatics software platform for visualizing molecular interaction networks and biological pathways and integrating these networks with annotations, gene expression profiles and other state data.
Although Cytoscape was originally designed for biological research, now it is a general platform for complex network analysis and visualization.
::DEVELOPER
:: SCREENSHOTS
:: REQUIREMENTS
- Windows / Linux / Mac OsX
- Java
:: DOWNLOAD
:: MORE INFORMATION
Citation:
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T.
Cytoscape: a software environment for integrated models of biomolecular interaction networks.
Genome Research 2003 Nov; 13(11):2498-504