ChemGenome 2.1
:: DESCRIPTION
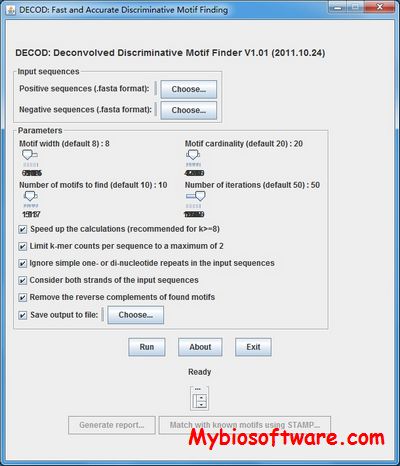
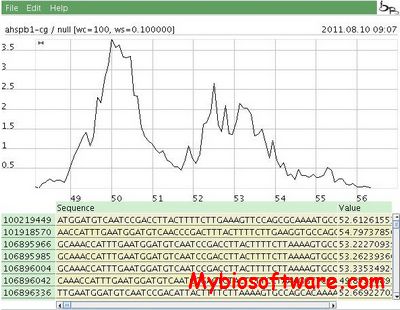
Chemgenome is an ab-intio gene prediction software, which find genes in prokaryotic genomes in all six reading frames. The methodology follows a physico-chemical approach and has been validated on 372 prokaryotic genomes.
::DEVELOPER
Supercomputing Facility for Bioinformatics & Computational Biology, IIT Delhi
:: SCREENSHOTS
N/A
:: REQUIREMENTS
- Linux
:: DOWNLOAD
:: MORE INFORMATION
Citation:
PLoS One. 2010 Aug 26;5(8):e12433.
A phenomenological model for predicting melting temperatures of DNA sequences.
Khandelwal G, Bhyravabhotla J.