BayeScan 2.1
:: DESCRIPTION
BayeScan ( (BAYEsian genome SCAN for outliers) ) aims at identifying candidate loci under natural selection from genetic data, using differences in allele frequencies between populations. BayeScan is based on the multinomial-Dirichlet model. One of the scenarios covered consists of an island model in which subpopulation allele frequencies are correlated through a common migrant gene pool from which they differ in varying degrees. The difference in allele frequency between this common gene pool and each subpopulation is measured by a subpopulation specific FST coefficient. Therefore, this formulation can consider realistic ecological scenarios where the effective size and the immigration rate may differ among subpopulations
::DEVELOPER
Computational and Molecular Population Genetics Lab, University of Bern
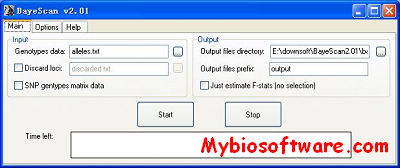
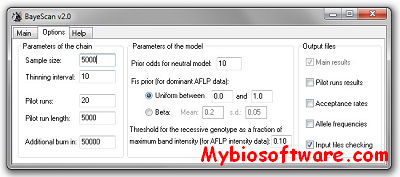
:: SCREENSHOTS
:: REQUIREMENTS
- Windows / Linux / Mac OsX
- R-Package
:: DOWNLOAD
:: MORE INFORMATION
Citation
Mol Ecol. 2011 Apr;20(7):1450-62. doi: 10.1111/j.1365-294X.2011.05015.x.
Enhanced AFLP genome scans detect local adaptation in high-altitude populations of a small rodent (Microtus arvalis).
Fischer MC, Foll M, Excoffier L, Heckel G.