Asterias
:: DESCRIPTION
Asterias is an integrated collection of freely-accessible web tools for the analysis of gene expression and aCGH data. Asterias includes applications for the analysis of genomic (and, to a lesser extent, proteomic) data that cover from data normalization to development of prediction models for survival data. Most of our applications use parallelization (via MPI) and run on a server with 60 CPUs for computation. And most of our applications allow the user to obtain additional information about the genes (chromosomal location, PubMed ids, Gene Ontology terms, etc) by using clickable links in tables and/or figures, thus allowing for enhanced interpretability of the results.
The tools include: normalization of expression and aCGH data (DNMAD); converting between different types of gene/clone and protein identifiers (IDconverter/IDClight); filtering and imputation (preP); finding differentially expressed genes related to patient class and survival data (Pomelo II); searching for models of class prediction (Tnasas); using random forests to search for minimal models for class prediction or for large subsets of genes with predictive capacity (GeneSrF); searching for molecular signatures and predictive genes with survival data (SignS); detecting regions of genomic DNA gain or loss (ADaCGH).
::DEVELOPER
Bioinfo Unit@CNIO
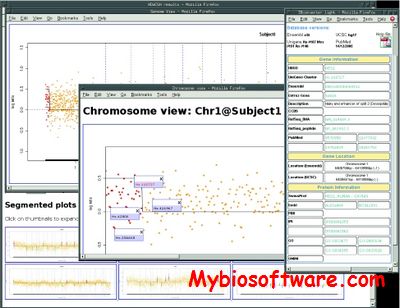
:: SCREENSHOTS
:: REQUIREMENTS
:: DOWNLOAD
:: MORE INFORMATION
Citation
Cancer Inform. 2007 Feb 3;3:1-9.
Asterias: a parallelized web-based suite for the analysis of expression and aCGH data.
Alibés A, Morrissey ER, Ca?ada A, Rueda OM, Casado D, Yankilevich P, Díaz-Uriarte R.